Editor’s Note: Today’s post is by Daniel S. Katz and Hollydawn Murray. Dan leads the FORCE11 Software Citation Implementation Working Group’s journals task force, and is the Chief Scientist at NCSA and Research Associate Professor in Computer Science, Electrical and Computer Engineering, and the School of Information Sciences at the University of Illinois. Until recently, Holly was the Head of Data and Software Publishing at F1000Research where she led on data and software sharing policy and strategy. She currently works as Research Manager at Health Data Research UK. The work discussed here is supported by representatives and editors from: AAS Journals, AGU Journals, American Meteorological Society, Crossref, DataCite, eLife, Elsevier, F1000Research, GigaScience Press, Hindawi, IEEE Publications, Journal of Open Research Software (Ubiquity Press), Journal of Open Source Software (Open Journals), Oxford University Press, PLOS, Science Magazine, Springer Nature, Taylor & Francis, and Wiley
Software is essential to research, and is regularly an element of the work described in scholarly articles. However, these articles often don’t properly cite the software, leading to problems finding and accessing it, which in turns leads to problems with reproducibility, reuse, and proper credit for the software’s developers. In response, the FORCE11 Software Citation Implementation Working Group, comprised of scholarly communications researchers, representatives of nineteen major journals, publishers, and scholarly infrastructures (Crossref, DataCite), have proposed a set of customizable guidelines to clearly identify the software and credit its developers and maintainers. This follows the earlier development of a set of Software Citation Principles. To realize their full benefit, we are now urging publishers to adapt and adopt these guidelines to implement the principles and to meet their communities’ particular needs.
Reproducibility
In March 2020, Fergusson et al. published a report that used software to model the impact of non-pharmaceutical interventions in reducing COVID-19 mortality and healthcare demand. This report, and others like it, were widely discussed and appeared to have had a quick impact on government policies. But almost as quickly, both political and scientific criticism started. The scientific criticism included questions about the modeling, such as the parameters and methodology used, which could not be easily addressed since the code was not shared in conjunction with the publication.
In 1995, Buckheit and Donoho paraphrased Claerbout and Karrenbach by saying, “an article about computational science in a scientific publication is not the scholarship itself, it is merely advertising of the scholarship. The actual scholarship is the full software development environment and the complete set of instructions which generated the figures.” Ideally this is readable source code, but in some cases may be an executable program or service. This software must be archived and cited properly so that it can be accessed and used to reproduce results.
Ferguson’s code was released in May, after a good deal of effort from Ferguson’s group, Microsoft, GitHub, and others. While this code is still subject to a great deal of debate, the debate has become more scientific, based on the code itself rather than assumptions about it.
Software reuse
In 2015, Topalidou & Rougier wanted to replicate, study, and reuse a model. While a paper using the model had been published, the code was not publicly available, and the paper was incomplete and partly inaccurate. The team did get a copy of the code, but found it overly complex and hard to use. They took three months to replicate it in Python, refine the model, make new predictions that were confirmed, and publish the reimplemented model (though without a license), which has now been used by others, as shown in Figure 1, as well as being further developed. This work was one of the inspirations for the ReScience Initiative. Similarly, Christopher Madan wanted to use software described by King et al., but he was unable to get access to it, so he rebuilt it as a toolbox, used it to address a science problem, and made it available on GitHub (with a license). This has now been reused and cited by others, for example by Canna et al. and Liu et al.
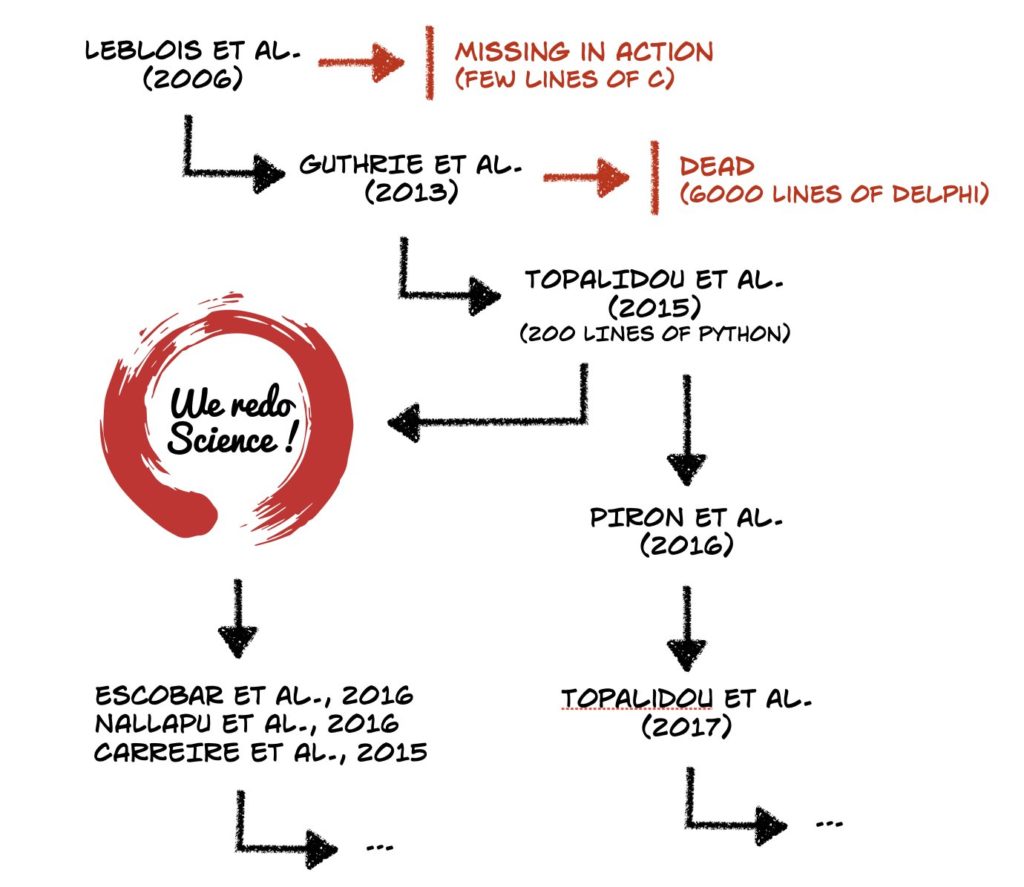
As Newton said in the context of science, “…if I have seen further, it is by standing on the shoulders of giants,” and using someone else’s software is a direct means to do this. The first image of a black hole was created using this idea — the researchers used existing instruments, including both hardware and software, then filled in the gaps to build on them to enable their discovery. This is the ideal we want to achieve: a researcher who wants to do new science can easily find and use existing software, and when needed, add to it and adapt it to meet their particular challenge. Having that existing software identifiable and retrievable from published papers is a step towards that ideal.
Software credit
Even when software work is widely recognized, the breadth of its application can be hard to measure if it is not cited. Previous research (e.g., Howison & Bullard) shows that software is typically not cited, though it is often mentioned in papers, sometimes in the methods sections. And when software is mentioned or cited, it happens in a number of inconsistent ways, including just stating the name of the software, placing a link to the software repository inline in the text, citing the user manual, citing a paper about the software, etc.
One example where software work was not recognized involves Travis Oliphant, an assistant professor at BYU in the early 2000s, who was the primary creator of NumPy, the Python library for numerical arrays that is the foundation of modern data science tools, and a primary creator of SciPy, the widely used library for scientific Python. These contributions weren’t sufficient for him to gain tenure in 2007, though there was enough curiosity about the potential impact of this work that he was granted an exception to the standard denial procedure and allowed to re-apply in two years. Travis left academia at this point, which was probably the right decision because it took until at least 2010 before the full impact of NumPy and SciPy was reasonably clear and until 2020 when Nature papers acknowledging the general impact of SciPy and NumPy were published. Of course, there are many more stories to be found from unhappy software developers, often including the fact that their software was used but not cited (See Figure 2).
So why isn’t citing a paper about the software good enough? A 2014 Nature article on the 100 most cited papers includes many papers about software, but even for these papers, they are a single point in time and a single set of authors: additional authors who work on the software after the publication are not credited, because the software itself (and the specific version that was used) is very seldom what is cited. For example, a 2012 paper on the Fiji image processing software has been cited over 20,000 times according to Google Scholar. Comparing the authors of this paper to the comments in the software’s GitHub repository makes it clear that some contributors are not getting credited by the papers citing it.
Sometimes software is cited and despite the numerous barriers that prevent people from receiving recognition and career success for their research software work, some have successfully overcome them. Dr. Fernando Perez, currently an associate professor in statistics at the University of California, Berkeley, is an example. For much of his career he worked in a traditional, untenured position as a research scientist in neuroscience and computational research, while collaboratively developing an open source notebook interface for the Python programming language as a side project. Over time, his software work started having much more of an impact than any of his traditional scientific contributions. The current evolution of his group’s efforts, the Jupyter ecosystem, is becoming widely accepted and used across the scholarly community and has led to awards such as the Association for Computer Machinery award for software. The magnitude of his software contributions and the far reaching impact of this effort earned him a fast-tracked tenured position at University of California, Berkeley.
Philanthropic funders including Wellcome, the Sloan Foundation, and the Chan Zuckerberg Initiative (CZI) and government agencies such as the NSF, NIH, UKRI, DFG, NASA, and more now recognize the importance of research software maintenance, growth, development, and community engagement via dedicated grant calls. For example, CZI has provided support to 72 open source tools through its Essential Open Source Software for Science program in the last two years. Matplotlib, an open-source Python plotting library, is one of many CZI grant recipients that embodies the discrepancy between software citation and impact. While preserved source code for Matplotlib has been cited 51 times since its deposition in Zenodo in 2016, the paper describing Matplotlib published in 2007 has been cited over 4,000 times (according to ADS). Nearly 300,000 other software packages depend on it. It’s worth noting that while dependencies will not necessarily always translate into citations, the number of Matplotlib dependent packages is on-par with the most cited scientific papers of all time. This, however, represents a further challenge faced by publishers: the muddied waters between citing the paper proposing the method and citing the source code itself. Not only does what is cited have implications for credit (as described earlier) but also for reproducibility.
Conclusions: Calling all publishers!
While software is increasingly essential to research, is increasingly funded as an independent project, and is often described in research papers that use it, practices for its formal citation have not kept up. Many publishers, including those who helped establish the guidelines inspiring this post, are already taking steps to ensure that the benefits of software citation are realized – and we urge you to join them! For example, Science’s editorial policies currently ask authors to cite the software central to their findings, and they plan to update this guidance with a link to the new article, The American Meteorological Society is in the process of developing a specific software policy and author guidelines that will be in place in 2021. Now is the time to engage editors and production teams, and educate editorial boards, authors, and reviewers. Along with the guidelines, we’re here to help publishers more broadly make software citation a consistent norm. And we’re now also starting to examine the next step in the publishing process: how software citations are captured, encoded, and transferred to the indexing and linking services provided by Crossref, DataCite, and others, where software information is currently being lost.
Discussion
4 Thoughts on "Guest Post — Citing Software in Scholarly Publishing to Improve Reproducibility, Reuse, and Credit"
Always a good idea to cite one’s sources, but why would software need a different citation format? The guidelines in the linked article look like APA format. Use whatever bibliographical standard is accepted in your field.
Software tools are not always citable by “normal APA” means. Who is the author and what is the year of publication of Adobe photoshop, or MATLAB, some of the most prevalent software tools in the scientific literature. I think that the goal is clear, but the implementation will not be straightforward. Dan I still think that RRIDs have a role to play here, they take care of the use case of citation a little better than APA citations for some software tools and authors don’t seem to mind them. I wish that software authors would be willing to add an entry to the RRID registry allowing us to track all the papers that mention their tools by name, or better yet RRID.
I agree that using established citations formats would be best. However, they don’t all have a format for software. We use Chicago Manual of Style, which does not have a software reference format. I found this FAQ that recommends not citing computer programs: https://www.chicagomanualofstyle.org/qanda/data/faq/topics/Documentation/faq0196.html
It would be helpful if APA, MLA, CMS, etc. developed reference formats for software. Then copyeditors, publishers, and programs like Mendeley and EndNote would be able to follow a standard format based on the reference guides already in use.
Great article and completely agree. Using these guidelines with automation can really help take this to a practical level. Code Ocean authors can create a standardize ‘compute capsule’ that is open, exportable, reproducible, and interoperable. Then with automation each capsule packages code, data, environment, and the associated results and is versioned. Authors automatically receive a DOI for the capsule and it can be embedded as a standard widget within an article.



